The capsid of the hepatitis B virus has a very unique structure. It is composed of many monomers which interact to provide flexibility to its structure and function.
Skip to:
- The hepatitis B virus
- Initiation of a hepatitis B viral infection
- The structure of the hepatitis B virus
- The effect of HAP1 on hepatitis B capsids
- Capsid-directed antiviral strategies
The hepatitis B virus
Hepatitis B virus (HBV) is a pararetrovirus that infects hepatocytes. HBV has an easily available vaccine but no cure. Approximately 257 million individuals are infected with HBV globally, and nearly 900,000 die each year related to disease complications, including liver failure or hepatic cancer.
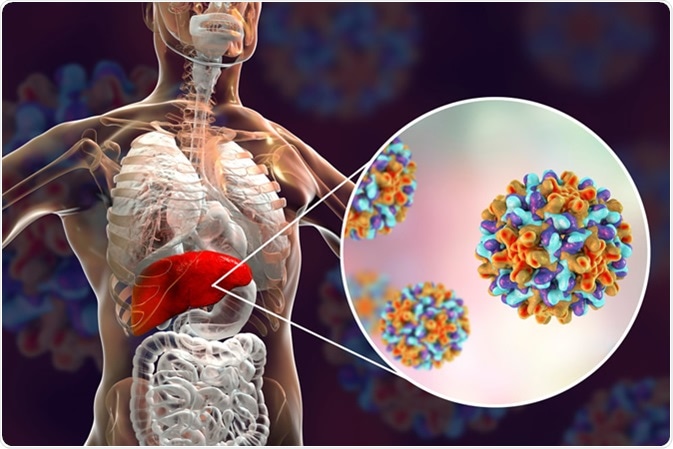
Initiation of a hepatitis B viral infection
The HBV surface is composed of HBV surface antigen (HBsAg) particles. The HBV virions (also referred to as Dane particles) are comprised of HBsAg particles. To initiate an infection, the Dane particle bind to sodium-taurocholate co-transporter polypeptides (NCTPs) on the surface of hepatocytes, which allows the virion to be transported into the cytoplasm of the hepatocytes via endocytosis. Following this the DNA filled core bind to α-importin β complexes on the core protein (HBcAg) surrounding the DNA, which allows the genome to be released. The HBV DNA is released as covalently closed circular DNA (cccDNA) which functions like host DNA, therefore, is less likely to be recognized as foreign.
An infected cell produces new HBV by making RNA copies of the HBV cccDNA. This produces copies of all the required proteins, antigens, and DNA. The mature HBV capsid initiates certain signaling cascades that allow for its envelopment and secretion so it can generate more cccDNA and infect more cells.
The structure of the hepatitis B virus
The hepatitis B virus is interesting in its ability to use relatively few virus proteins to carry out a wide range of actions. HBcAg exists as a soluble dimer with an RNA binding domain. The structure of HBcAg dimers can vary with dimers in capsids being more compact than free dimers in solution. Regulation of HBcAg can change the conformational state of the dimers which affects their functional state. An example being the oxidization of core protein dimers which leads to weaker dimer-dimer interactions, slowing down capsid assembly. The ability for HBcAg to switch between different structural states is crucial for its function in capsid assembly and HBV virology.
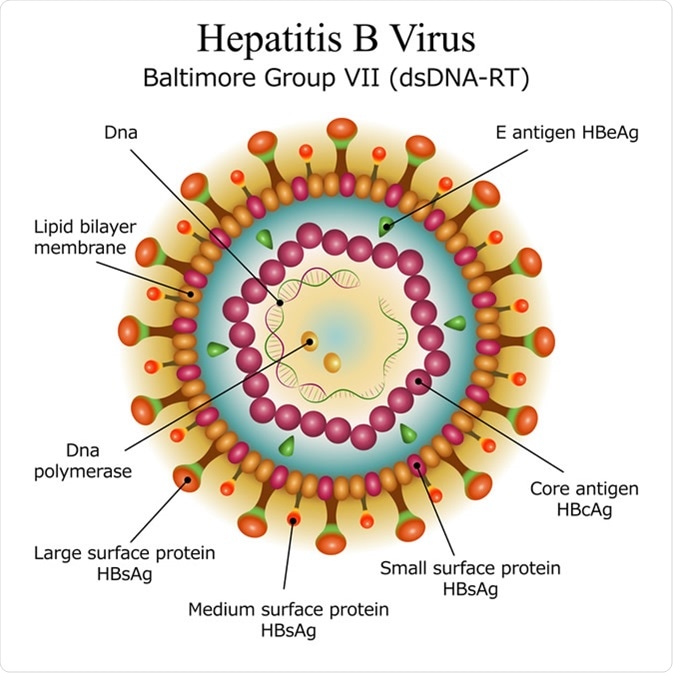
HBeAg (a variant of the HBcAg) is a secreted protein expressed by HBV. HBeAg has 2 main functional parts. The first is made up of 19 amino acid residues and is responsible for transporting the protein to the endoplasmic reticulum. The other part is made of 10 residues and binds to HBcAg via an intramolecular disulfide bond which stabilizes both HBcAg and HBeAg. HBeAg is important for the immune evasion properties of HBV.
As previously mentioned, HBcAg assembles to form the virus capsid. The capsid is an icosahedron composed of 120 copies of capsid protein homodimer. Each T=4 unit is composed of 4 HBcAg monomers (two dimers; AB and CD). The HBcAg capsid is extremely immunogenic, so induces both B-cell and T-cell immune responses; however, evidence suggests that those responses are not protective against an HBV infection. Due to its versatility and immunogenicity, HBcAg capsids can be used as carriers for epitopes in vaccines.
The effect of HAP1 on hepatitis B capsids
A heteroarylidihydropyrimidine (HAP1) is an HBV capsid assembly activator and misdirector. Knowledge of this activity would be extremely beneficial to the development of antiviral strategies against HBV infections. The capsid structure has been analyzed with and without the influence of HAP1. This analysis discovered that HAP1 causes structural changes to capsids by moving capsid subunits, leading to an altered icosahedral structure. These changes are initiated by HAP1 binding sites on capsid monomers. These binding sites can also interfere with the capsid protein-protein interactions which cause4s changes to the capsid quaternary structure leading to misdirections in assembly. This shows that HAP1 can be both constructive and destructive to the formation of HBV capsids.
HAP1 molecules used on cultured cells leads to the decreased production of virions and the increased loss of capsid proteins via proteasomal digestion. HAP1 was shown to accelerate capsid assembly but at stoichiometric concentrations, it misdirects capsid assembly, which correlates with the expected outcome, and shows the flexibility of capsid functions. Additionally, HAP1 can be bound to existing capsids to trigger disassembly. Future research in this area hopes to utilize the HAP1 binding site on capsid proteins to create anti-HBV treatments.
Capsid-directed antiviral strategies
HBcAg can potentially be used as an antiviral target due to its role in the formation of capsids and the life cycle of HBV. Also, as capsids have no human homolog, they make great targets for therapeutic intervention. Research has shown that altering temperature and ionic strength of the HBcAg monomers can lead to the formation of aberrant non-capsid structures and trapped intermediates, which causes disruption to the capsid assembly. Taking advantage of this may be an effective antiviral method to combat HBV infections.
Targeting HBV virology may be achieved using non-nucleoside reverse transcriptase inhibitors; phenyl propenamides (PPAs) and Heteroaryldihysropyrimidines (HAPs). Using PPAs on HBV leads to the loss of cytoplasmic RNA-filled cores and the formation of empty capsids. Using HAPs leads to the loss of core proteins in HBV. Both PPAs and HAPs achieve their effects by speeding up the assembly of protein-protein interactions. In high enough concentrations of PPA or HAP, this leads to non-icosahedral capsid complexes and the incorrect assembly of core proteins, which is severely detrimental to the correct development and functions of HBV. Both of these reverse transcriptase inhibitors also strengthen dimer-dimer bond associations, effectively doubling the rate of assembly, which can further disrupt the capsid assembly.
Sources
- Venkatakrishnan, B. and Zlotnick, A. (2016). The Structural Biology of Hepatitis B Virus: Form and Function. Annual Review of Virology. https://dx.doi.org/10.1146%2Fannurev-virology-110615-042238
- Hadden, J., et al. (2018). All-atom molecular dynamics of the HBV capsid reveals insights into biological function and cryo-EM resolution limits. ELife. https://doi.org/10.7554/eLife.32478.001
- Bourne, C., et al. (2006). Global Structural Changes in Hepatitis B Virus Capsids Induced by the Assembly Effector HAP1. Journal of Virology. https://jvi.asm.org/content/80/22/11055
Further Reading
- All Hepatitis B Content
- What is Hepatitis B?
- Hepatitis B Mechanisms
- Hepatitis B Diagnosis
- Hepatitis B Prevention
Last Updated: Aug 20, 2019
Written by
Samuel Mckenzie
Sam graduated from the University of Manchester with a B.Sc. (Hons) in Biomedical Sciences. He has experience in a wide range of life science topics, including; Biochemistry, Molecular Biology, Anatomy and Physiology, Developmental Biology, Cell Biology, Immunology, Neurology and Genetics.
Source: Read Full Article
